If you’ve been paying attention to COVID lately, you will know that there is a new variant on the scene. It’s called BA.2.86, and it’s a bit surprising: essentially it comes from a slightly unexpected part of the omicron evolutionary tree, and has various mutations which suggest that it might grow fast. This is coloured by our experience of the original BA.1 omicron, which also branched off from an unexpected place, had various mutations, and really did grow fast - doubling around every two days (40% or higher daily growth rate) in the UK for example.
So it’s really important that we get an estimate of the BA.2.86 growth rate. In this post, I’m going to give you an estimate, and then explain why you should ignore it. In doing so, I want to pay tribute to the great Indian statistician C.R. Rao, whose death at the age of 102 was announced today. Rao was a remarkable figure who bridges many generations of statistics, from his PhD under Ronald Fisher right through to his co-authorship of a textbook on neural networks published in his 100th year. So what can Rao tell us about BA.2.86?
The key thing (as so often) is the idea of a statistical model. So here’s a simple one. We expect exponential growth (or decay!) of this variant, so we’ll imagine that in week x the proportion of cases made up of this variant will be c times rho to the power x, or c rho^x for short. Here, rho represents the weekly growth rate - if rho was 2, then you’d come back a week later and find twice as many cases as now.
Now, each week we take a different number of tests for sequencing - say the number in week x is n(x). What we’d expect is that the number of tests u(x) that come back as BA.2.86 would have what’s called a Poisson distribution with mean n(x) c rho^x. (Don’t worry about why! In fact, you can skip this paragraph if you like!)
The key thing is that we have a familiar framework: we have data (n(x), u(x)) and we have what statisticians call parameters, namely (c, rho). (The names of these parameters are chosen in deliberate tribute!). The game is this: we want to use the known data to get a good estimate of the unknown parameters.
One way that we can do this is using something called the likelihood. It’s a slightly odd beast: we essentially think about the probability that we would see the data for particular values of the parameters. Since the data is fixed, we can think of varying the parameters, and seeing which choice of parameters makes this probability biggest. This is called maximum likelihood estimation.
It’s a nice idea, and somewhat intuitive. We are trying to find the model under which the observed data was most likely - that is when is P(data given parameters) biggest? However, it’s important to note that we are not quite doing the thing we’d really like to do - that is finding the most likely parameters, by making P(parameters given data) biggest. (Though if you are a Bayesian statistician, you’ll know that the two things are somewhat related - the likelihood tells you the weighting you put on your prior beliefs when you update them).
The great news is that we can actually do this with BA.2.86. Thanks to people sequencing genomes worldwide, uploading them to GISAID, we have exactly the kind of weekly data that we need. We can plug this into the formula … and get an estimate of rho that equals 3.85 - corresponding to a daily growth rate of 21%, or 3.6 day doubling. That’s not great! It’s not as fast as original omicron BA.1, but it’s faster than the BA.5 advantage which caused a fair amount of trouble back in summer 2022.
So, this seems terrible. If I was unscrupulous, I could probably post this estimate to Twitter, stick in a couple of graphs and harvest a few hundred retweets. So why am I not doing so? And what has this got to do with C.R. Rao?
The issue is that I took an arbitrary decision to choose the parameters that make the likelihood biggest. But the likelihood itself isn’t a fixed thing: it depends on the data we saw, which we know is somewhat random (who gets tested? how fast are things reported?). So different data would have given us a different likelihood function.
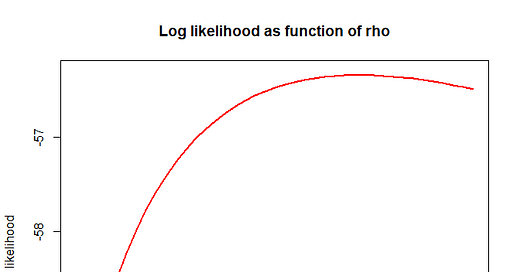
But a better way to see that my choice of 21% is somewhat arbitrary is just to plot a graph of the likelihood itself. In fact we plot the logarithm of the likelihood (this isn’t just me being my usual log-obsessed self, there are good reasons why the logarithm is a good thing to use - it often makes the calculation easier, and it means that combining independent data makes the logarithm of the likelihoods add up). So here goes:
What you’ll see is that the log-likelihood is pretty flat. While in general we believe that big likelihoods are associated with the right parameters, it’s pretty close to a tie at the top. Although the value of the likelihood at our “best guess” 3.85 is higher than the value at say 2 or 5 (a much tamer 10% daily growth rate or an even worse 26% rate), it’s not exactly a slam dunk: a different random sample of data would have brought a different curve. Indeed, it’s possible to argue with our model - was the data really sampled at random, were reports of BA.2.86 expedited on discovery?
In other words our estimate of rho is somewhat uncertain, there’s a range of plausible values, and we shouldn’t be putting guesses out on Twitter for clicks. And certainly it indicates the dangers of just relying on a mechanistic procedure (“find where the biggest value is”) without at least thinking about what this graph might look like.
This is really where Rao’s work comes in. In general, the log-likelihood will look like this kind of parabola (“tennis ball throws”, as I describe them in Numbercrunch). We can measure how confident we should be about the estimate by thinking about how pointy the parabola is. If, like here, it’s flattish (doesn’t curve much) then we aren’t very confident about our estimate. On the other hand if it was very thin and pointy (straight up and down) then there are only a few parameter choices compatible with the data and it seems much more likely that our estimate is true.
One of Rao’s great contributions was to quantify this intuition, in terms of a quantity called the Fisher information. This measures the bendiness (curvature) of the parabola: flat parabolas have low Fisher information, narrow ones have high Fisher information. The result known as the Cramér–Rao lower bound tells us that if a model has low Fisher information (like in the BA.2.86 case), then it’s not possible to estimate our parameters precisely - however smart we are.
This is pretty stunning stuff. It turns out there are fundamental limits to estimation, and they arise through thinking about the geometry (the curviness) of the likelihood function. Indeed, Rao opened up links between statistics and geometry in this way, and essentially founded the field of information geometry by thinking about surfaces in space formed by the parameters of the statistical model.
Rao made many other great contributions in the fundamental theory of estimation, such as the Rao-Blackwell theorem which he proved together with the great statistician David Blackwell, and which gives a recipe to find the optimal estimator in a statistical problem. And as I have tried to illustrate here, his ideas are still extremely relevant to key problems today.
It may seem like the position is hopeless: we have an estimate, but we don’t know what to do with it. However, the usual bunny rules apply - we just need to wait! As we get more data, the likelihood will tend to become pointier, as the influence of any one reported number is reduced. So with time it should become much more clear what the true growth advantage of BA.2.86 really is - but in the meantime it’s worth remembering that the worldwide prevalence is currently likely to be at most a few per cent, so if it’s going to cause a wave (and it may well do), it’s probably not going to do it at this exact instant.




Very nice read - took me back statistics lectures in the 70s at London Uni - well some of it did, the rest was brand new - thank you Oliver
Great explanation of the likelihood phenomenon. Thanks! I may borrow some of this to support my estimates, if you don’t mind.